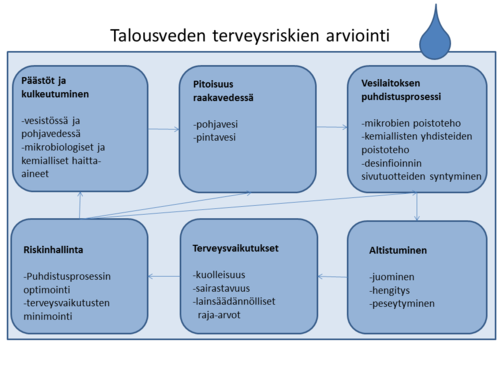
+ Näytä koodi- Piilota koodi
library(OpasnetUtils)
library(ggplot2)
library(reshape2)
if(RaakaLuokka != 'Käyttäjä') {
Kampylo <- 'Luokituksen mukainen'
Ecoli <- 'Luokituksen mukainen'
Rota <- 'Luokituksen mukainen'
Noro <- 'Luokituksen mukainen'
Crypto <- 'Luokituksen mukainen'
Giardia <- 'Luokituksen mukainen'
}
IRaakaPitoisuus <- list(Kampylo, Ecoli, Rota, Noro, Crypto, Giardia)
# Vedenkulutus
if (!is.na(suppressWarnings(as.numeric(IVedenkulutus)))) {
InpVedenkulutus = data.frame(VedenkulutusResult = IVedenkulutus)
InpVedenkulutus = Ovariable("Vedenkulutus", output = InpVedenkulutus)
}
# Patogeenien pitoisuudet
objects.latest("Op_fi2655", "muuttuja") # [[Raakaveden patogeenipitoisuudet]] RaakaPitoisuus, customoitia varten
RaakaPitoisuus <- EvalOutput(RaakaPitoisuus)
temp <- RaakaPitoisuus@output
temp <- temp[as.character(temp$Raakavesilähde) == RaakaLuokka, ]
temp <- merge(
temp,
data.frame(
Patogeeni = c("Kampylobakteeri","E.coli O157:H7","Rotavirus","Norovirus","Cryptosporidium","Giardia"),
TempResult = suppressWarnings(as.numeric(IRaakaPitoisuus))
)
)
temp$RaakaPitoisuusResult <- ifelse(
is.na(temp$TempResult),
temp$RaakaPitoisuusResult,
temp$TempResult
)
RaakaPitoisuus@output <- temp[, !colnames(temp) %in% c("TempResult", "Raakavesilähde")]
temp <- rep(FALSE,9) # vaihtoehdot + klooraus joka on mallin rakenteesta johtuen aina epätosi
temp[Puhdistus] <- TRUE
Puhdistus <- temp
if (!is.na(suppressWarnings(as.numeric(IKlooriAnnos)))) if (IKlooriAnnos >= 0) {
InpKlooriAnnos <- Ovariable("KlooriAnnos", output = data.frame(KlooriAnnosResult = IKlooriAnnos))
}
# Käsitellään puhdistusmenetelmät käsin tätä mallia varten
riippuvuudet <- data.frame(
Name = c("RaakaPitoisuus", "Desinfiointi", "PuhdistusTeho", "KloorausTeho"),
Ident = c("Op_fi2655/muuttuja", "Op_fi2993/muuttuja", "Op_fi2656/muuttuja", "Op_fi2667/tehokkuus")
)
objects.latest("Op_fi2993", "muuttuja") # [[Juomaveden desinfiointi]] Desinfiointi: klooraus ym
objects.latest("Op_fi2656", "muuttuja") # [[Vedenkäsittelyn tehokkuus]] PuhdistusTeho: suodatukset ym
PuhdistusTeho <- EvalOutput(PuhdistusTeho)
Desinfiointi <- EvalOutput(Desinfiointi)
Puhdistus <- data.frame(
Käytössä = Puhdistus,
Puhdistusmenetelmä = c("Perinteinen puhdistus", "Hyvin toimva puhdistus",
"Tehostettu puhdistus" ,"Hidas hiekkasuodatus", "Kalkkikivisuodatus",
"Aktiivihiilisuodatus", "UV", "Otsonointi", "Klooraus")
)
PuhdistusTeho@output <- merge(PuhdistusTeho@output, Puhdistus[1:6,]) # Vedenkäsittelymenetelmät
Desinfiointi@output <- merge(Desinfiointi@output, Puhdistus[7:9,]) # Desinfiointimenetelmät
PuhdistusTeho@output$PuhdistusTehoResult <- ifelse(
PuhdistusTeho@output$Käytössä,
PuhdistusTeho@output$PuhdistusTehoResult,
0
)
Desinfiointi@output$DesinfiointiResult <- ifelse(
Desinfiointi@output$Käytössä,
Desinfiointi@output$DesinfiointiResult,
0
)
# Poistetaan "Käytössä" kolumni
PuhdistusTeho@output <- PuhdistusTeho@output[, !colnames(PuhdistusTeho@output) == "Käytössä"]
Desinfiointi@output <- Desinfiointi@output[, !colnames(Desinfiointi@output) == "Käytössä"]
objects.latest("Op_fi1759", "muuttuja") # [[Infektion todennäköisyys per henkilö]] Vaste
Vaste <- EvalOutput(Vaste, substitute = TRUE)
#################################################################################
dose.response = Vaste@output
Pathogen <- c("Kampylobakteeri","E.coli O157:H7","Rotavirus","Norovirus","Cryptosporidium","Giardia")
vaesto <- opbase.data("Op_fi2652")[,c("Ikä","Result")] # [[Suomen ikäjakauma]]
colnames(vaesto) <- c("Age", "Osuus")
vaesto$Populaatio <- vaesto$Osuus * Vaestonkoko
odotettu.elinika <- 81
colnames(dose.response)[colnames(dose.response) == "Patogeeni"] <- "Pathogen"
colnames(dose.response)[colnames(dose.response) == "VasteResult"] <- "P.inf"
colnames(dose.response)[colnames(dose.response) == "AltistuminenResult"] <- "Exp.pat"
P.ill.g.inf <- data.frame(Pathogen, P.ill.g.inf = c(0.33, 1 - (270 / 1540), 0.9, 0.7, 0.71, 1)) # todennäköisyys sairastua kun saa infektion
# Kampylobakteeri, DALYt per infektio
P.treat.g.ill.Kamp.Gastr <- data.frame(Pathogen = Pathogen[c(1,1,1)], Outcome = "Gastroenteritis", ill.treat = c("Untreated",
"General practitioner", "Hospitalised", "Unspecified")[c(1,2,3)], P.treat.g.ill = c(0.7627, 0.2373, 0.0097))
P.treat.ill.g.inf.Kamp.Gastr <- merge(P.treat.g.ill.Kamp.Gastr, P.ill.g.inf)
P.treat.ill.g.inf.Kamp.Gastr$P.treat.ill.g.inf <- P.treat.ill.g.inf.Kamp.Gastr$P.ill.g.inf *
P.treat.ill.g.inf.Kamp.Gastr$P.treat.g.ill
duration.ill.treat.Kamp.Gastr <- data.frame(Outcome = c("Gastroenteritis"), ill.treat = c("Untreated", "General practitioner",
"Hospitalised", "Unspecified")[c(1,2,3)], dur.ill = c(5.1 / 365, 8.4 / 365, 14.39 / 365))
severity.ill.treat.Kamp.Gastr <- data.frame(Outcome = c("Gastroenteritis"), ill.treat = c("Untreated", "General practitioner",
"Hospitalised", "Unspecified")[c(1,2,3)], sev.ill = c(0.067, 0.393, 0.393))
daly.ill.treat.Kamp.Gastr <- merge(P.treat.ill.g.inf.Kamp.Gastr, duration.ill.treat.Kamp.Gastr)
daly.ill.treat.Kamp.Gastr <- merge(daly.ill.treat.Kamp.Gastr, severity.ill.treat.Kamp.Gastr)
daly.ill.treat.Kamp.Gastr$dalys <- daly.ill.treat.Kamp.Gastr$P.treat.ill.g.inf * daly.ill.treat.Kamp.Gastr$dur.ill *
daly.ill.treat.Kamp.Gastr$sev.ill
P.death.g.ill.Gastr <- 0.0004
P.death.g.inf.Gastr <- P.death.g.ill.Gastr * P.ill.g.inf$P.ill.g.inf[P.ill.g.inf$Pathogen == "Kampylobakteeri"]
death.Gastr.life.lost <- 13.2
daly.death.Kamp.Gastr <- P.death.g.inf.Gastr * death.Gastr.life.lost
## GBS Kamp.
P.gbs.g.ill <- 2e-004
P.gbs.g.inf <- P.gbs.g.ill * P.ill.g.inf$P.ill.g.inf[P.ill.g.inf$Pathogen == "Kampylobakteeri"]
dur.sev.factor.gbs <- data.frame(Outcome = c("Clinical GBS", "Residual GBS"), dur.sev.factor = c(0.29, 5.8)) # duration * severity * fraction?
daly.Kamp.gbs <- data.frame(dur.sev.factor.gbs$Outcome, dalys = dur.sev.factor.gbs$dur.sev.factor * P.gbs.g.inf)
P.death.g.gbs <- 0.08 / 3 # triangular 0.01, 0.02, 0.05
P.death.g.inf.gbs <- P.death.g.gbs * P.gbs.g.inf
death.gbs.life.lost <- 18.7
daly.death.Kamp.gbs <- P.death.g.inf.gbs * death.gbs.life.lost
## reactive arthritis Kamp.
P.arth.g.ill <- 0.02 # triangluar 0.01, 0.02, 0.03
P.arth.g.inf <- P.arth.g.ill * P.ill.g.inf$P.ill.g.inf[P.ill.g.inf$Pathogen == "Kampylobakteeri"]
duration.arth <- 6 / 52
severity.arth <- 0.21
daly.Kamp.arth <- P.arth.g.inf * duration.arth * severity.arth
# E.coli
P.wd.g.ill <- 0.53 # watery diarrhea
P.wd.g.inf <- P.wd.g.ill * P.ill.g.inf$P.ill.g.inf[P.ill.g.inf$Pathogen == "E.coli O157:H7"]
severity.wd <- 0.067
duration.wd <- 3.4 / 365
daly.wd.Ecoli <- P.wd.g.inf * severity.wd * duration.wd
P.hc.g.ill <- 0.47
P.hc.g.inf <- P.hc.g.ill * P.ill.g.inf$P.ill.g.inf[P.ill.g.inf$Pathogen == "E.coli O157:H7"]
severity.hc <- 0.39
duration.hc <- 5.6 / 365
daly.hc.Ecoli <- P.hc.g.inf * severity.hc * duration.hc
P.death.g.ill.Ecoli <- 0.00027
P.death.g.inf.Ecoli <- P.death.g.ill.Ecoli * P.ill.g.inf$P.ill.g.inf[P.ill.g.inf$Pathogen == "E.coli O157:H7"]
age.death.Ecoli <- 81 - 13.2
daly.death.Ecoli <- P.death.g.inf.Ecoli * (odotettu.elinika - age.death.Ecoli)
## Haemolytic uraemic syndrome (HUS)
P.hus.g.ill <- 0.01
P.hus.g.inf <- P.hus.g.ill * P.ill.g.inf$P.ill.g.inf[P.ill.g.inf$Pathogen == "E.coli O157:H7"]
severity.hus <- 0.93
duration.hus <- 21 / 365
daly.hus.Ecoli <- P.hus.g.inf * severity.hus * duration.hus
P.death.g.hus <- 0.04
P.death.hus.g.inf <- P.death.g.hus * P.hus.g.inf
age.death.hus.Ecoli <- 81 - 26.2
daly.death.hus.Ecoli <- P.death.hus.g.inf * (odotettu.elinika - age.death.hus.Ecoli)
## End Stage Renal Disease (ESRD)
P.esrd.g.hus <- 0.118
P.esrd.g.inf <- P.hus.g.inf * P.esrd.g.hus
severity.duration.hus <- 8.7 # severity * duration
daly.esrd.Ecoli <- P.esrd.g.inf * severity.duration.hus
P.death.g.esrd <- 0.0252
P.death.esrd.g.inf <- P.esrd.g.inf * P.death.g.esrd
age.death.esrd.Ecoli <- 81 - 34
daly.death.esrd.Ecoli <- P.death.esrd.g.inf * (odotettu.elinika - age.death.esrd.Ecoli)
# Rotavirus
P.treat.g.ill.Rotavirus <- data.frame(Pathogen = "Rotavirus", ill.treat = c("Untreated",
"General practitioner", "Hospitalised")[rep(1:3, each = 82)], Age = rep(0:81, 3), P.treat.g.ill = c(rep(0.82,5),
rep(0.95, 10), rep(0.99, 50), rep(0.97, 17), rep(0.137, 5), rep(0.0244, 5), rep(0.0511, 5), rep(0.0127, 50),
rep(0.0299, 17), rep(0.0416, 5), rep(0.0213, 5), rep(0, 72)))
P.treat.ill.g.inf.Rotavirus <- merge(P.treat.g.ill.Rotavirus, P.ill.g.inf)
P.treat.ill.g.inf.Rotavirus$P.treat.g.inf <- P.treat.ill.g.inf.Rotavirus$P.ill.g.inf * P.treat.ill.g.inf.Rotavirus$P.treat.g.ill
duration.ill.treat.Rotavirus <- data.frame(ill.treat = c("Untreated", "General practitioner","Hospitalised"), dur.ill = c(4.9 / 365,
7.1 / 365, 7.7 / 365))
severity.ill.treat.Rotavirus <- data.frame(ill.treat = c("Untreated", "General practitioner", "Hospitalised"), sev.ill = c(0.067,
0.393, 0.393))
daly.ill.treat.Rotavirus <- merge(P.treat.ill.g.inf.Rotavirus, duration.ill.treat.Rotavirus)
daly.ill.treat.Rotavirus <- merge(daly.ill.treat.Rotavirus, severity.ill.treat.Rotavirus)
daly.ill.treat.Rotavirus$dalys <- daly.ill.treat.Rotavirus$P.treat.g.inf * daly.ill.treat.Rotavirus$dur.ill *
daly.ill.treat.Rotavirus$sev.ill
P.death.Rotavirus <- data.frame(Age = 0:81, P.death.g.ill = c(rep(2.13e-005, 5), rep(0, 77)))
P.death.Rotavirus$P.death.g.inf <- P.death.Rotavirus$P.death.g.ill * P.ill.g.inf$P.ill.g.inf[P.ill.g.inf$Pathogen == "Rotavirus"]
P.death.Rotavirus$Life.lost <- odotettu.elinika - P.death.Rotavirus$Age
daly.death.Rotavirus <- data.frame(Age = P.death.Rotavirus$Age, dalys = P.death.Rotavirus$P.death.g.inf * P.death.Rotavirus$Life.lost)
# Norovirus
P.treat.g.ill.Norovirus <- data.frame(Pathogen = "Norovirus", ill.treat = c("Untreated",
"General practitioner", "Hospitalised")[rep(1:3, each = 82)], Age = rep(0:81, 3), P.treat.g.ill = c(rep(0.94876706,5),
rep(0.9902, 5), rep(0.98239, 5), rep(0.98434, 51), rep(0.992741, 16), rep(0.0448,5), rep(8.6e-003, 5), rep(0.0154, 5),
rep(0.0137, 51), rep(6.17e-003, 16), rep(6.43e-003,5), rep(1.2e-003, 5), rep(2.21e-003, 5), rep(1.96e-003, 51),
rep(8.85e-004, 16)))
P.treat.ill.g.inf.Norovirus <- merge(P.treat.g.ill.Norovirus, P.ill.g.inf)
P.treat.ill.g.inf.Norovirus$P.treat.g.inf <- P.treat.ill.g.inf.Norovirus$P.ill.g.inf * P.treat.ill.g.inf.Norovirus$P.treat.g.ill
duration.ill.treat.Norovirus <- data.frame(ill.treat = c("Untreated", "General practitioner","Hospitalised"), dur.ill = c(3.8 / 365,
5.73 / 365, 7.23 / 365))
severity.ill.treat.Norovirus <- data.frame(ill.treat = c("Untreated", "General practitioner", "Hospitalised"), sev.ill = c(0.067,
0.393, 0.393))
daly.ill.treat.Norovirus <- merge(P.treat.ill.g.inf.Norovirus, duration.ill.treat.Norovirus)
daly.ill.treat.Norovirus <- merge(daly.ill.treat.Norovirus, severity.ill.treat.Norovirus)
daly.ill.treat.Norovirus$dalys <- daly.ill.treat.Norovirus$P.treat.g.inf * daly.ill.treat.Norovirus$dur.ill *
daly.ill.treat.Norovirus$sev.ill
P.death.Norovirus <- data.frame(Age = 0:81, P.death.g.ill = c(rep(2.94e-006, 5), rep(0, 61), rep(2.04e-004, 16)))
P.death.Norovirus$P.death.g.inf <- P.death.Norovirus$P.death.g.ill * P.ill.g.inf$P.ill.g.inf[P.ill.g.inf$Pathogen == "Norovirus"]
P.death.Norovirus$Life.lost <- odotettu.elinika - P.death.Norovirus$Age
daly.death.Norovirus <- data.frame(Age = P.death.Norovirus$Age, dalys = P.death.Norovirus$P.death.g.inf * P.death.Norovirus$Life.lost)
# Cryptosporidium
P.treat.g.ill.Crypt <- data.frame(Pathogen = "Cryptosporidium", ill.treat = c("Untreated",
"General practitioner", "Hospitalised")[rep(1:3, each = 82)], Age = rep(0:81, 3), P.treat.g.ill = c(rep(0.9175730049999999,5),
rep(0.80937, 5), rep(0.6810499999999999, 5), rep(0.9774191, 50), rep(0.94706, 17), rep(0.082,5), rep(0.188, 5), rep(0.316, 5),
rep(0.0209, 50), rep(0.0367, 17), rep(4.26e-004,5), rep(2.63e-003, 5), rep(2.95e-003, 5), rep(1.66e-003, 50), rep(0.0146, 17)))
P.treat.ill.g.inf.Crypt <- merge(P.treat.g.ill.Crypt, P.ill.g.inf)
P.treat.ill.g.inf.Crypt$P.treat.g.inf <- P.treat.ill.g.inf.Crypt$P.ill.g.inf * P.treat.ill.g.inf.Crypt$P.treat.g.ill
duration.ill.treat.Crypt <- data.frame(ill.treat = c("Untreated", "General practitioner","Hospitalised"), dur.ill = c(3.5 / 365,
7 /365, 18.4 / 365))
severity.ill.treat.Crypt <- data.frame(ill.treat = c("Untreated", "General practitioner", "Hospitalised"), sev.ill = c(0.067,
0.393, 0.393))
daly.ill.treat.Crypt <- merge(P.treat.ill.g.inf.Crypt, duration.ill.treat.Crypt)
daly.ill.treat.Crypt <- merge(daly.ill.treat.Crypt, severity.ill.treat.Crypt)
daly.ill.treat.Crypt$dalys <- daly.ill.treat.Crypt$P.treat.g.inf * daly.ill.treat.Crypt$dur.ill *
daly.ill.treat.Crypt$sev.ill
P.death.Crypt <- data.frame(Age = 0:81, P.death.g.ill = c(rep(9.95e-007, 5), rep(0, 10), rep(2.09e-005, 50), rep(1.64e-003, 17)))
P.death.Crypt$P.death.g.inf <- P.death.Crypt$P.death.g.ill * P.ill.g.inf$P.ill.g.inf[P.ill.g.inf$Pathogen == "Cryptosporidium"]
P.death.Crypt$Life.lost <- odotettu.elinika - P.death.Crypt$Age
daly.death.Crypt <- data.frame(Age = P.death.Crypt$Age, dalys = P.death.Crypt$P.death.g.inf * P.death.Crypt$Life.lost)
# Giardia
P.treat.g.ill.Giardia <- data.frame(Pathogen = "Giardia", ill.treat = c("Untreated",
"General practitioner", "Hospitalised")[rep(1:3, each = 82)], Age = rep(0:81, 3), P.treat.g.ill = c(rep(0.9376,5),
rep(0.91034, 5), rep(0.72642, 5), rep(0.92486, 50), 0.54596, rep(0.5365, 16), rep(0.0609,5), rep(0.0852, 5), rep(0.272, 5),
rep(0.0721, 50), rep(0.451, 17), rep(1.5e-003,5), rep(4.46e-003, 5), rep(1.58e-003, 5), rep(3.04e-003, 51), rep(0.0125, 16)))
P.treat.ill.g.inf.Giardia <- merge(P.treat.g.ill.Giardia, P.ill.g.inf)
P.treat.ill.g.inf.Giardia$P.treat.g.inf <- P.treat.ill.g.inf.Giardia$P.ill.g.inf * P.treat.ill.g.inf.Giardia$P.treat.g.ill
duration.ill.treat.Giardia <- data.frame(ill.treat = c("Untreated", "General practitioner","Hospitalised"), dur.ill = c(10 / 365,
10 /365, 30 / 365))
severity.ill.treat.Giardia <- data.frame(ill.treat = c("Untreated", "General practitioner", "Hospitalised"), sev.ill = c(0.067,
0.393, 0.393))
daly.ill.treat.Giardia <- merge(P.treat.ill.g.inf.Giardia, duration.ill.treat.Giardia)
daly.ill.treat.Giardia <- merge(daly.ill.treat.Giardia, severity.ill.treat.Giardia)
daly.ill.treat.Giardia$dalys <- daly.ill.treat.Giardia$P.treat.g.inf * daly.ill.treat.Giardia$dur.ill *
daly.ill.treat.Giardia$sev.ill
# yhteenveto DALYistä
Health.effects <- vaesto[,c("Age","Populaatio")]
Health.effects$Untreated.Gastr.Kamp <- daly.ill.treat.Kamp.Gastr[daly.ill.treat.Kamp.Gastr$ill.treat == "Untreated", c("dalys")]
Health.effects$GP.Gastr.Kamp <- daly.ill.treat.Kamp.Gastr[daly.ill.treat.Kamp.Gastr$ill.treat == "General practitioner", c("dalys")]
Health.effects$Hospitalised.Gastr.Kamp <- daly.ill.treat.Kamp.Gastr[daly.ill.treat.Kamp.Gastr$ill.treat == "Hospitalised", c("dalys")]
Health.effects$Death.Gastr.Kamp <- daly.death.Kamp.Gastr
Health.effects$Clinical.GBS.Kamp <- daly.Kamp.gbs$dalys[1]
Health.effects$Residual.GBS.Kamp <- daly.Kamp.gbs$dalys[2]
Health.effects$Death.GBS.Kamp <- daly.death.Kamp.gbs
Health.effects$Arth.Kamp <- daly.Kamp.arth
Health.effects$WD.Ecoli <- daly.wd.Ecoli
Health.effects$HC.Ecoli <- daly.hc.Ecoli
Health.effects$Death.Ecoli <- daly.death.Ecoli
Health.effects$HUS.Ecoli <- daly.hus.Ecoli
Health.effects$Death.HUS.Ecoli <- daly.death.hus.Ecoli
Health.effects$ESRD.Ecoli <- daly.esrd.Ecoli
Health.effects$Death.ESRD.Ecoli <- daly.death.esrd.Ecoli
Health.effects <- merge(Health.effects, daly.ill.treat.Rotavirus[daly.ill.treat.Rotavirus$ill.treat == "Untreated", c("Age", "dalys")])
colnames(Health.effects)[ncol(Health.effects)] <- "Untreated.Rotavirus"
Health.effects <- merge(Health.effects, daly.ill.treat.Rotavirus[daly.ill.treat.Rotavirus$ill.treat == "General practitioner", c("Age", "dalys")])
colnames(Health.effects)[ncol(Health.effects)] <- "GP.Rotavirus"
Health.effects <- merge(Health.effects, daly.ill.treat.Rotavirus[daly.ill.treat.Rotavirus$ill.treat == "Hospitalised", c("Age", "dalys")])
colnames(Health.effects)[ncol(Health.effects)] <- "Hospitalised.Rotavirus"
Health.effects <- merge(Health.effects, daly.death.Rotavirus)
colnames(Health.effects)[ncol(Health.effects)] <- "Death.Rotavirus"
Health.effects <- merge(Health.effects, daly.ill.treat.Norovirus[daly.ill.treat.Norovirus$ill.treat == "Untreated", c("Age", "dalys")])
colnames(Health.effects)[ncol(Health.effects)] <- "Untreated.Norovirus"
Health.effects <- merge(Health.effects, daly.ill.treat.Norovirus[daly.ill.treat.Norovirus$ill.treat == "General practitioner", c("Age", "dalys")])
colnames(Health.effects)[ncol(Health.effects)] <- "GP.Norovirus"
Health.effects <- merge(Health.effects, daly.ill.treat.Norovirus[daly.ill.treat.Norovirus$ill.treat == "Hospitalised", c("Age", "dalys")])
colnames(Health.effects)[ncol(Health.effects)] <- "Hospitalised.Norovirus"
Health.effects <- merge(Health.effects, daly.death.Norovirus)
colnames(Health.effects)[ncol(Health.effects)] <- "Death.Norovirus"
Health.effects <- merge(Health.effects, daly.ill.treat.Crypt[daly.ill.treat.Crypt$ill.treat == "Untreated", c("Age", "dalys")])
colnames(Health.effects)[ncol(Health.effects)] <- "Untreated.Crypt"
Health.effects <- merge(Health.effects, daly.ill.treat.Crypt[daly.ill.treat.Crypt$ill.treat == "General practitioner", c("Age", "dalys")])
colnames(Health.effects)[ncol(Health.effects)] <- "GP.Crypt"
Health.effects <- merge(Health.effects, daly.ill.treat.Crypt[daly.ill.treat.Crypt$ill.treat == "Hospitalised", c("Age", "dalys")])
colnames(Health.effects)[ncol(Health.effects)] <- "Hospitalised.Crypt"
Health.effects <- merge(Health.effects, daly.death.Crypt)
colnames(Health.effects)[ncol(Health.effects)] <- "Death.Crypt"
Health.effects <- merge(Health.effects, daly.ill.treat.Giardia[daly.ill.treat.Giardia$ill.treat == "Untreated", c("Age", "dalys")])
colnames(Health.effects)[ncol(Health.effects)] <- "Untreated.Giardia"
Health.effects <- merge(Health.effects, daly.ill.treat.Giardia[daly.ill.treat.Giardia$ill.treat == "General practitioner", c("Age", "dalys")])
colnames(Health.effects)[ncol(Health.effects)] <- "GP.Giardia"
Health.effects <- merge(Health.effects, daly.ill.treat.Giardia[daly.ill.treat.Giardia$ill.treat == "Hospitalised", c("Age", "dalys")])
colnames(Health.effects)[ncol(Health.effects)] <- "Hospitalised.Giardia"
Health.effects <- reshape(Health.effects, idvar = c("Age"), times = colnames(Health.effects)[-c(1,2)], timevar = "Outcome",
varying = list(colnames(Health.effects)[-c(1,2)]), direction = "long")
colnames(Health.effects)[4] <- "P.daly.g.inf"
Health.effects$Pathogen <- NA
Health.effects$Pathogen[grep(".Kamp", Health.effects$Outcome)] <- Pathogen[1]
Health.effects$Pathogen[grep(".Ecoli", Health.effects$Outcome)] <- Pathogen[2]
Health.effects$Pathogen[grep(".Rotavirus", Health.effects$Outcome)] <- Pathogen[3]
Health.effects$Pathogen[grep(".Norovirus", Health.effects$Outcome)] <- Pathogen[4]
Health.effects$Pathogen[grep(".Crypt", Health.effects$Outcome)] <- Pathogen[5]
Health.effects$Pathogen[grep(".Giardia", Health.effects$Outcome)] <- Pathogen[6]
Health.effects <- merge(Health.effects, dose.response[,c("Pathogen", "P.inf")])
Health.effects$DALYs <- (1 - (1 - Health.effects$P.inf * Health.effects$P.daly.g.inf)^365) * Health.effects$Populaatio
temp <- merge(dose.response, P.ill.g.inf)
############# EXTRA ###########################################
MikrobienLogVähenemä <- combine(Desinfiointi, PuhdistusTeho, KloorausTeho, name = "MikrobienLogVähenemä")
MikrobienLogVähenemä@output[["Puhdistusmenetelmä"]][
MikrobienLogVähenemä@output[["MikrobienLogVähenemäSource"]] == "KloorausTehoFormula"
] <- "Klooraus"
#MikrobienLogVähenemä <- oapply(MikrobienLogVähenemä, cols = c("Puhdistusmenetelmä", "KlooriAnnosSource"), FUN = sum, na.rm = TRUE)
#MikrobienLogVähenemä@output <- fillna(MikrobienLogVähenemä@output, colnames(MikrobienLogVähenemä@output)[MikrobienLogVähenemä@marginal])
############# TULOKSET #########################################################################################################
cat("Patogeenien konsentraatio raakavedessä\n")
oprint(RaakaPitoisuus) # Patogeenien konsentraatio raakavedessä
cat("Patogeenien log vähenemä puhdistuksessa (Kuva 1.)\n")
ggplot(MikrobienLogVähenemä@output, aes(x = Patogeeni, weight = MikrobienLogVähenemäResult, fill = Puhdistusmenetelmä)) +
geom_bar() + theme_gray(base_size = 24) + labs(title = "Kuva 1. Mikrobien log-vähenemä")
cat("Patogeeneille altistuminen ja infektion todennäköisyys\n")
oprint(Vaste@output[,c("Patogeeni", "AltistuminenResult", "VasteResult")]) # Patogeeneille altistuminen ja infektion todennäköisyys (vaste)
cat("Arvioitu terveysvaikutus\n")
cat(sum((1 - (1 - temp$P.ill.g.inf * temp$P.inf)^365) * Vaestonkoko, na.rm = TRUE), " vatsatautia vuodessa \n")
cat(sum(Health.effects$DALYs, na.rm = TRUE), " DALY:ä vatsataudeista \n")
| |